Paraview video5/11/2023 
Many of the ParaView examples use Show() Render(), which generates a frozen, non-interactive window. ParaView users can quickly build visualizations to analyze their data using qualitative and quantitative. Note that the frames are rendered as fast as possible. The number of frames is controlled by the No. In Sequence mode, the animation is played as a sequence of images (or frames) generated one after the other and rendered in immediate succession. If your computer has a discrete GPU, 1024 or more points per dimension may be possible, with proportionately slower rendering.Ī key feature of ParaView is that it gracefully degrades to render 3D output even on modest devices, as long as you don't select too large Sampling Dimensions. ParaView supports three modes for playing animation. If your computer does not have a discrete GPU, using more than about 256 grid points in a dimension may crash ParaView, even with 16 GB RAM. The data can be sampled with Property "Sampling Dimensions" at higher resolution, but this takes proportionately more GPU resources. Then, in the Properties browser, select Representation: Volume. nc filename, Add Filter, Resample as Image In the "Pipeline Browser" right click the. The data needs to be filtered to display as a volume in ParaView. Large datasets no longer need to be downloaded and exchanged, and colleagues can get instantaneous feedback as Omniverse users can work in the same workspace in the cloud. because ipympl is not properly installed, you can as a. and for Jupyter Lab use this instead: matplotlib widget. 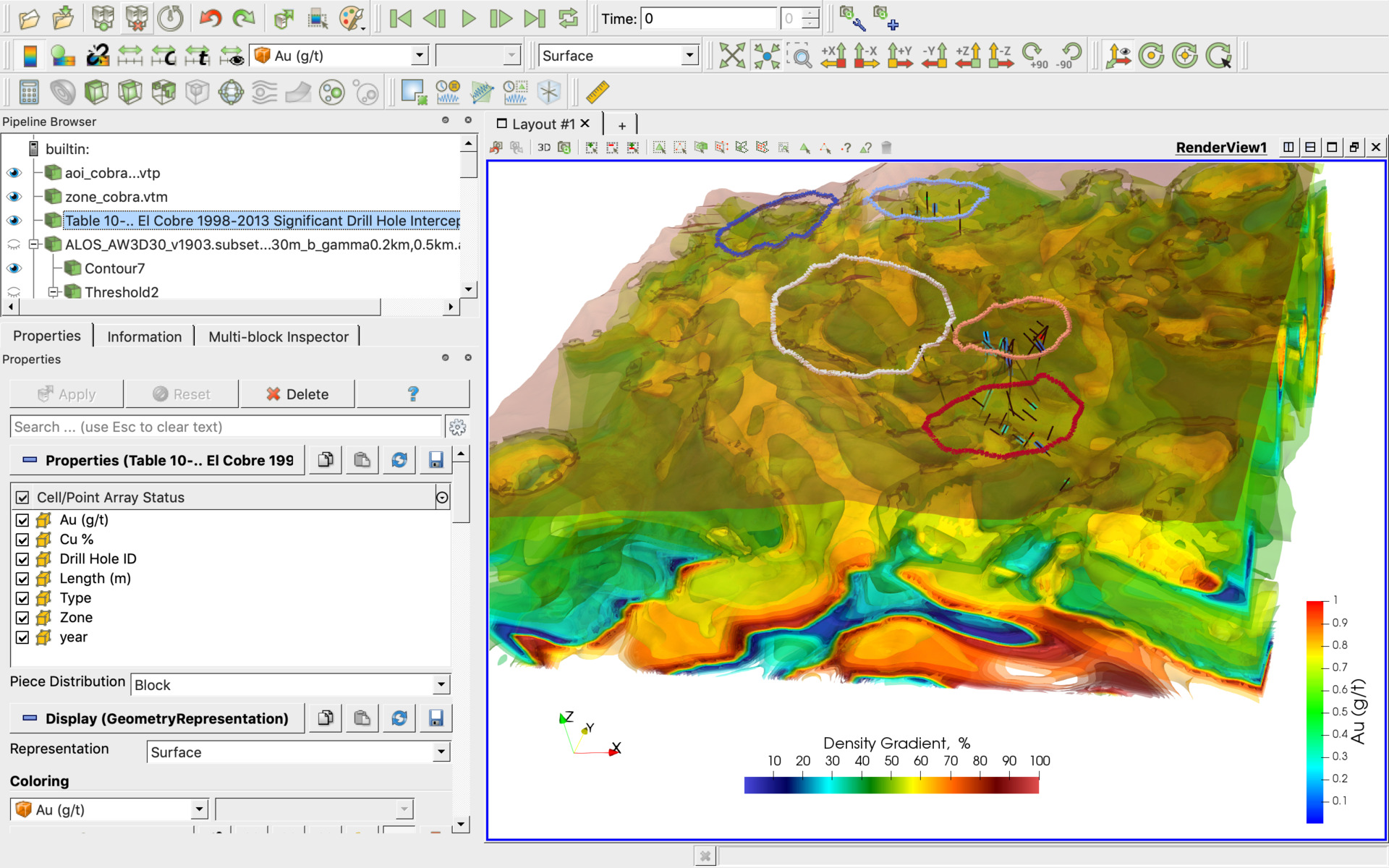
For interactive plots in Jupyter notebook, add this 'cell magic' to the first line of your notebook: matplotlib notebook. To save animations in pvpython, you use SaveAnimation. openPMD-viewer can be used either in simple Python scripts or in Jupyter. If saving as images, ParaView will generate a series of image files sequentially numbered using the frame number as a suffix to the specified filename. The "convert_data" scripts store the grid in the NetCDF file. Now available by downloading Omniverse Open Beta, the Omniverse ParaView Connector enables researchers to boost their productivity and speed their discoveries. The available file formats include AVI and Ogg (when available) video formats, as well as image formats such PNG, JPEG, and TIFF. To use a grid, which is particularly important for non-uniform gridded ionospheric data, store the NetCDF4 data with the grid data embedded in the file. ParaView can load single-variable NetCDF4 files. In the ParaView GUI, select "Volume" and the variable name to actually render the data. However, non-COARDS, non-CF single variable files can be loaded with the plain NetCDF filter. As a consequence, some parts of the webinar or its entirety may not be applicable to. Windows Gemini users are already using MS-MPI, so they should also use the Windows ParaView MPI installer. Please be aware that this webinar was developed for our legacy systems. MacOS and Linux have only MPI ParaView downloads. The example using cuml "UMAP MNIST Example.ipynb" requires a Linux system with GPU due toĬuml is a general machine learning GPU library unrelated to IPyParaView.Īlso the Dask example "Dask-MPI_Volume_Render" is for running on Dask-MPI HPC cluster, which would need to be setup by your IT staff perhaps. 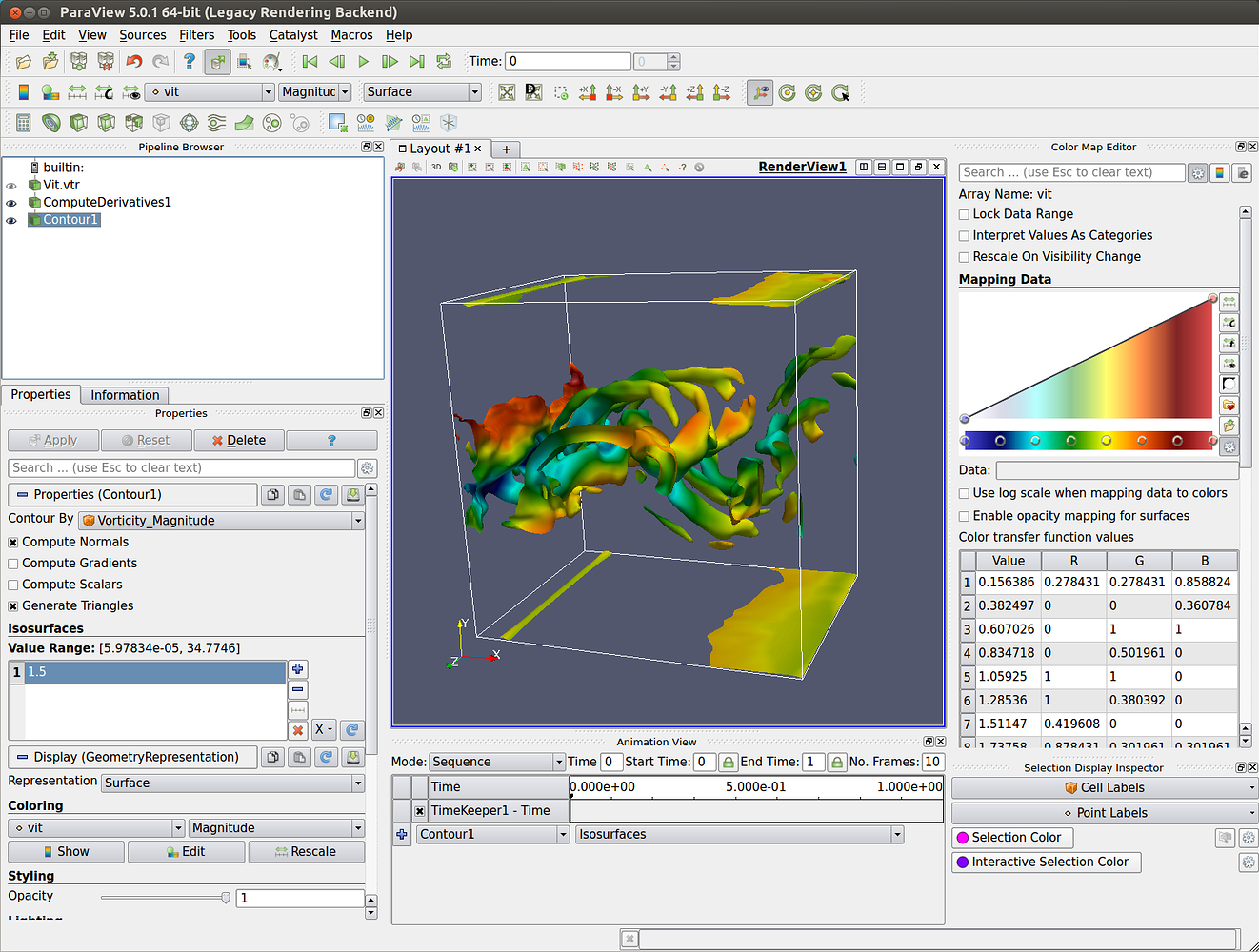
The data exploration can be done interactively. 2 It can be used to build visualizations to analyze data using qualitative and quantitative techniques. ParaView is known and used in many different communities to analyze and visualize scientific data sets. Iso-Surfaces_with_RTX.ipynb "RTX" requires an appropriate GPUĪdvanced examples (not for beginning user) ParaView is an open-source, multi-platform data analysis and visualization application.We suggest examples below, which do not require a discrete GPU (they run on a light laptop): Hope that helps - I’m curious to hear what works best for your use case.And use the web browser Jupyter Notebook that automatically opens to browse and run the examples. That’s why the client-side rendering is preferred in general, although there will be limits on how much data can be downloaded efficiently and how much can be loaded into the browser. This actually works quite nicely, but it means you have to maintain a server with the user’s data loaded or create one on the fly. OHIF client for dicom study browser, layout, app contextįor server-side rendering with Slicer, I’ve been playing for a while with embedding a web server in Slicer.DICOMweb serving all the data (can come from Google Healthcare or self-hosted open source server).Maybe you could describe a bit about what you want to achieve and why your current approach isn’t working for you and why you want to change direction?įor myself, we’ve been working mostly lately with this formulation (all should be easy to find by searching but let me know if you want more info): I’m not personally too familiar with the paraview approaches. Sounds like you are exploring all the good options, but probably there isn’t yet a really complete end-to-end implementation that’s in production.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
